Principal Investigator
Other positions:
Director Graduate School of Medical Genetics, University of Campania “Luigi Vanvitelli", Naples, Italy
Professor of Medical Genetics, Department of Precision Medicine, University of Campania Luigi Vanvitelli, Italy
Sandro Banfi obtained his Medical Degree and undertook his Neurology Residency at the University of Naples “Federico II”. In 1991, he moved to the Department of Pediatrics at Baylor College of Medicine where he worked on a post-doctoral project aimed at the elucidation of the genetic basis of Spinocerebellar Ataxias. In 1995, he moved back to Italy where he joined TIGEM as an independent investigator and shifted his research focus to data mining and functional genomics approaches in order to identify biologically relevant genes with potential roles in disease pathogenesis. In 2019, he became Full Professor of Medical Genetics at University of Campania “Luigi Vanvitelli”. He is also coordinator of the PhD program in Human Genetics at TIGEM in affiliation with the UK Open University. Sandro Banfi has made significant contributions to the definition of the molecular basis of Inherited Retinal Diseases and to the identification and characterization of a number of genes, both coding and noncoding, endowed with relevant roles in retinal biology.
Study of the molecular basis of inherited eye disorders with a focus on the role of microRNAs
Inherited eye diseases are a common cause of visual impairment in children and young adults. They include both neurodegenerative forms, including, among others, retinitis pigmentosa (RP), Stargardt Disease and Leber Congenital Amaurosis (LCA), and developmental defects, such as microphthalmia, anophthalmia and coloboma. All these diseases are characterized by extensive genetic heterogeneity, which constitutes an obstacle both for accurate diagnosis and when devising effective gene-based treatments.
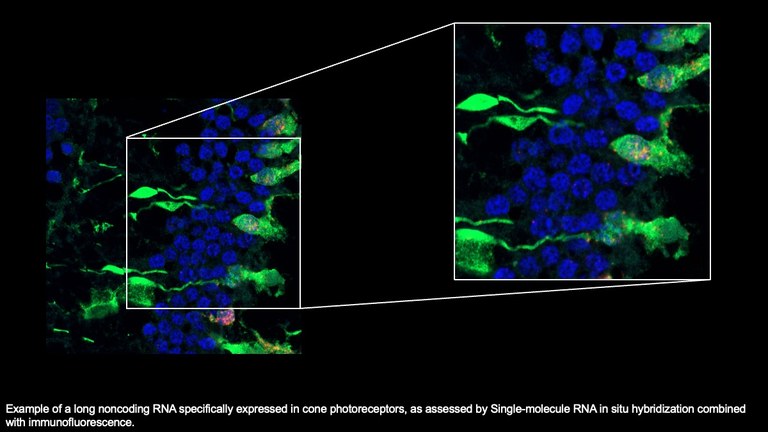
In the past 20 years, we have focused our efforts on the elucidation of the role of microRNAs and other non-coding RNAs in eye development and function, in both physiological and pathological conditions. In particular, we have generated the most comprehensive atlas of microRNA expression data (miRNome) in the eye available to date, first based on RNA in situ hybridization and subsequently on next generation sequencing (NGS) approaches. We have identified the first microRNA responsible for a monogenic form of ocular disease, namely miR-204, which is mutated in an autosomal dominant RP. We also demonstrated that the Adeno-Associated Viral (AAV) administration of miR-204 exerts a protective effect on photoreceptors in different models of inherited degenerations in a mutation/gene-independent manner. We are now extending this analysis to additional microRNAs that represent potential gene-agnostic therapeutic tools for inherited eye diseases. We believe that this strategy may represent a valid alternative/complementary approach to gene-based therapy and could be extended to a larger portion of patients in the near future. Finally, we also aim to shed light on the “missing heritability” of a significant fraction of patients with Inherited eye diseases who remain undiagnosed even after extensive NGS-based genomic approaches, such as targeted or whole exome sequencing using an integrated strategy aimed at the search for structural variants and single nucleotide variants with an impact on transcript integrity.
- In vitro high-content screening reveals miR-429 as a protective molecule in photoreceptor degeneration. Mol Ther Nucleic Acids. 2024
- miR-181a/b downregulation: a mutation-independent therapeutic approach for inherited retinal diseases. EMBO Molecular Medicine, 2022
- AAV-miR-204 Protects from Retinal Degeneration by Attenuation of Microglia Activation and Photoreceptor Cell Death. Molecular Therapy - Nucleic Acids, 2019
- High-resolution analysis of the human retina miRNome reveals isomiR variations and novel microRNAs. Nucleic Acids Research, 2016
- MiR-204 is responsible for inherited retinal dystrophy associated with ocular coloboma. PNAS, 2015
The full list of publications can be visualized in Google Scholar
Quote
My research goals are to elucidate the molecular basis of inherited disorders, and particularly of neurodegenerative conditions. I am now particularly interested in gaining insight into the contribution of noncoding RNAs to pathophysiological processes.
Additional Funding
FFB - FOUNDATION FIGHTING BLINDNESS - Pleiotropic effect of AAVmiRNA /sponge and mechanisms of protection (2024-2025)











